 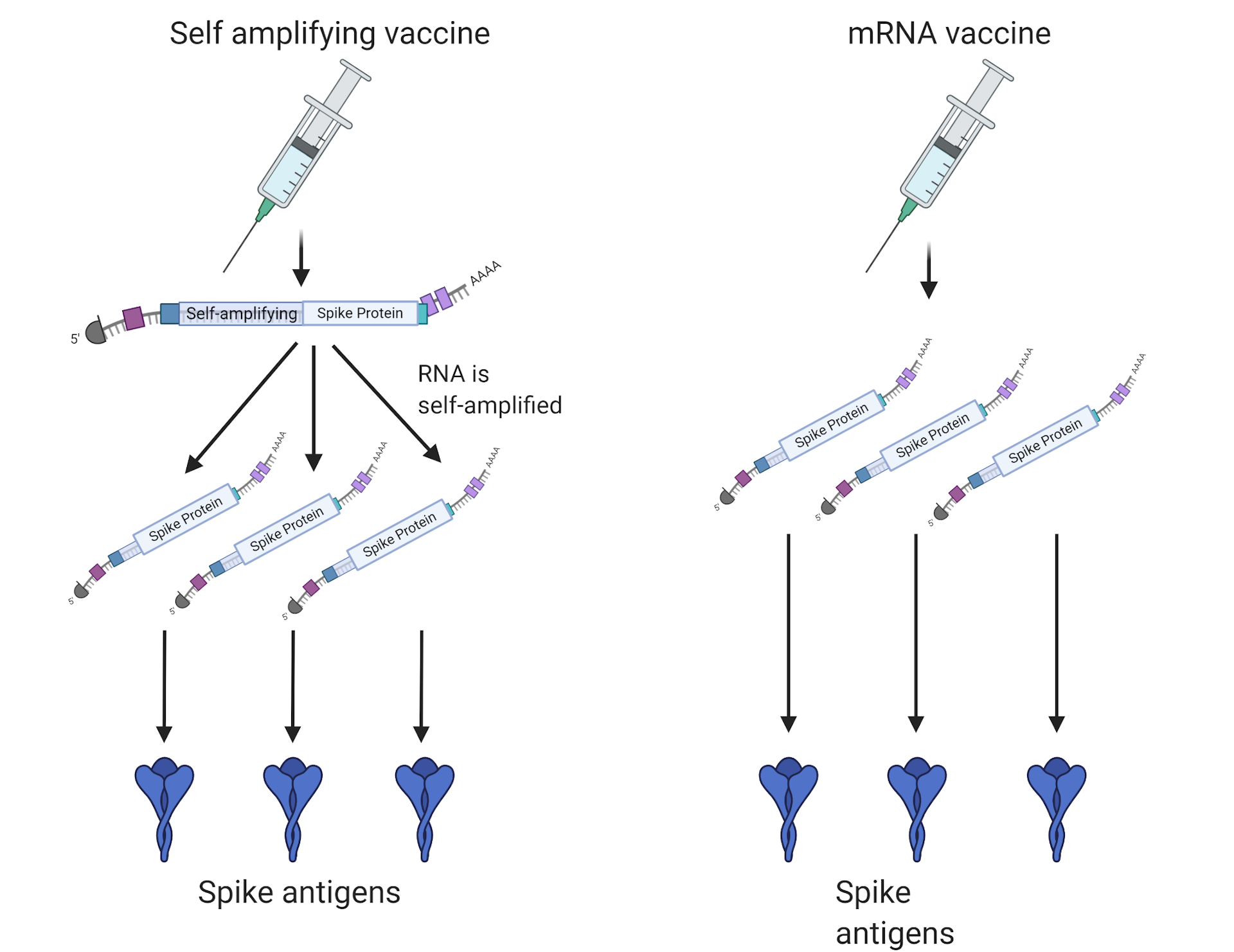
A relatively recent approach for vaccine development, first proposed by Coleman et al. Given the urgency to combat this pandemic, multiple efforts to develop effective vaccines are underway. As of Jthere have been 6,931,000 confirmed cases and 400,857 deaths from COVID-19 worldwide 1. The recent emergence of the 2019 novel coronavirus (SARS-CoV-2) has gained worldwide attention and sparked an international effort to develop treatments and vaccines. Our analysis identified the spike (S) and nucleocapsid (N) proteins as promising targets for deoptimization and suggests a roadmap for SARS-CoV-2 vaccine development, which can be generalizable to other viruses. We performed comprehensive in silico analyses of several features of SARS-CoV-2 genomic sequence (e.g., codon usage, codon pair usage, dinucleotide/junction dinucleotide usage, RNA structure around the frameshift region) in comparison with other members of the coronaviridae family of viruses, the overall human genome, and the transcriptome of specific human tissues such as lung, which are primarily targeted by the virus. Therefore, it is well suited for emerging viruses, for which we may not have extensive data. This approach carries the advantage that it only requires limited knowledge specific to the virus in question, other than its genome sequence. 
A promising approach for vaccine development is to generate, through codon pair deoptimization, an attenuated virus. As the SARS-CoV-2 pandemic is rapidly progressing, the need for the development of an effective vaccine is critical. 
0 Comments
Leave a Reply. |
 RSS Feed
RSS Feed
